2017
Andromeda
idea
In May, 2017, XAOP presented a Satellite Workshop at FlandersBio’s Life Sciences conference Knowledge for Growth. The workshop’s main themes combined Machine Learning, Amazon Web Services (AWS), and High Performance Computing.
The goal of this model was to find out how we could return other suggestions for related words without returning similar words, like PubMed currently returns its similar suggestions. By related we mean that they have something in common but without being too obvious - for example, searching for depression in PubMed gives us results like postpartum depression, major depression, ... while our model returns insomnia, MDD, anxiety, … The model we created gives users the advantage of being able to broaden their search terms with some context, instead of narrowing them.
approach
Using abstracts of PubMed ranging from 2010-2017 as our public data source, we set up a search engine that returns words related to the word searched for using the Word2Vec model. Word2Vec is a model that creates word embeddings from a corpus.
For our workshop, we used PubMed abstracts. When words are in a vector space, an interesting result occurs: given a word, the model returns words that are related (by using a distance metric, e.g. cosine similarity) thereby providing the user with some context.
technology
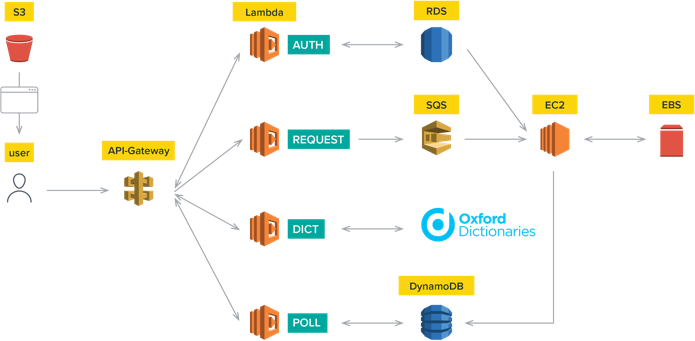
Everything is hosted on Amazon Web Services from the mining and processing to the learning and visualisations. It starts with fetching data from PubMed, then processing and formatting these data to fit our requirements. Word2Vec models are subsequently trained with different parameters where every model is then evaluated by mathematical equations and human testing. These models are all trained on the servers (EC2 instances) of AWS. The chosen model is combined with the Andromeda website where Andromeda sends words to the model and the model returns related words. These results are then visualised with a plot using the cosine similarity.
All the models are trained on the EC2 instances of AWS. Andromeda sends words to the Andromeda website to the chosen model and the model returns related words. These results are visualised with a plot using the cosine similarity.
services
other projects
ThirdEye
Development
Technology
Architecture
AML
Development
Technology
Architecture
Halcyon
Development
Design
Technology
SmartWithFood platform
Development
Support
Biocartis Diagnostics grid
Scientific business analysis
Development
Design
Xpectrum
Development
Design
Architecture
Cartographer
Scientific business analysis
Development
Design